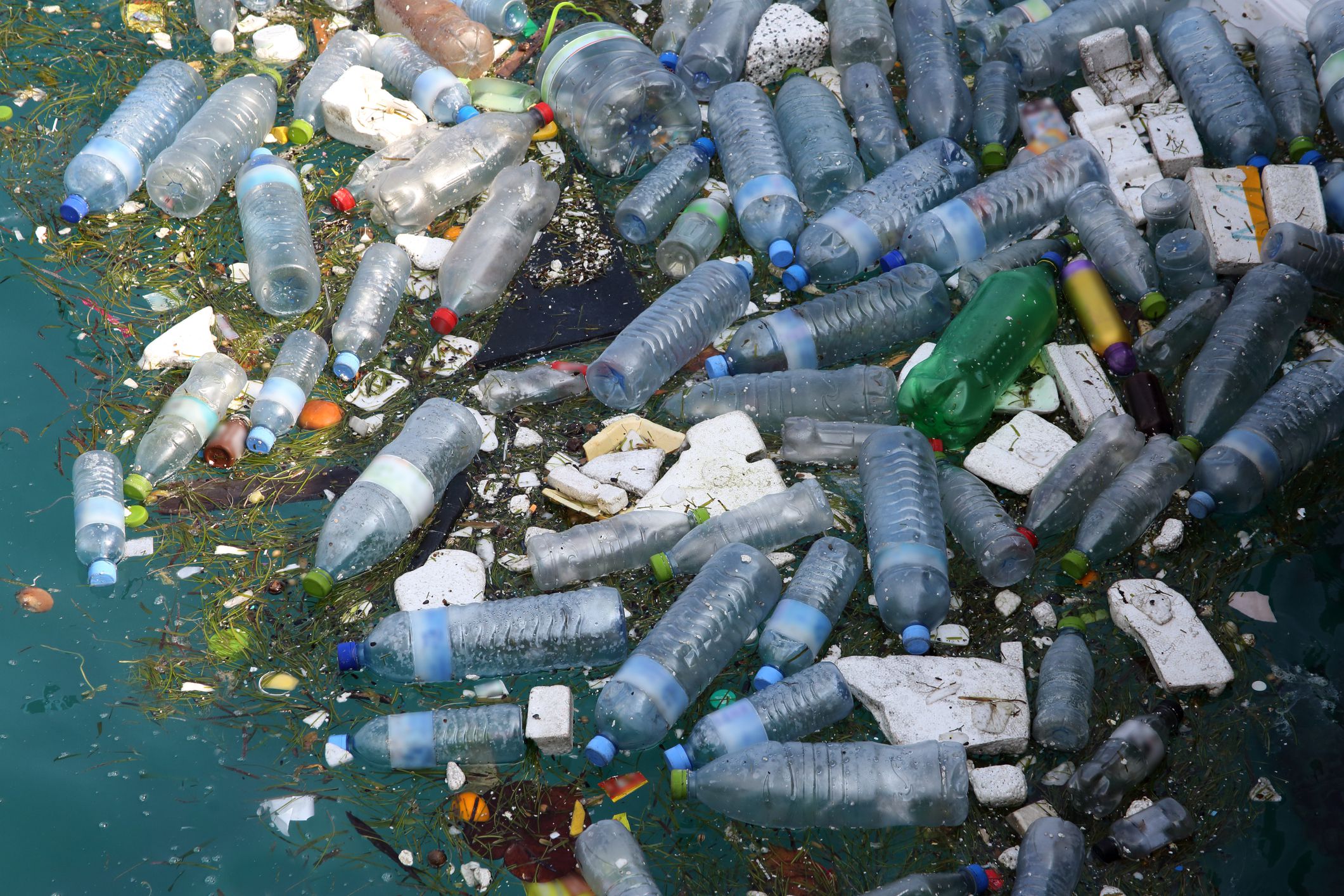
Researchers have discovered enzymes able to break down plastics far faster than was possible a decade ago
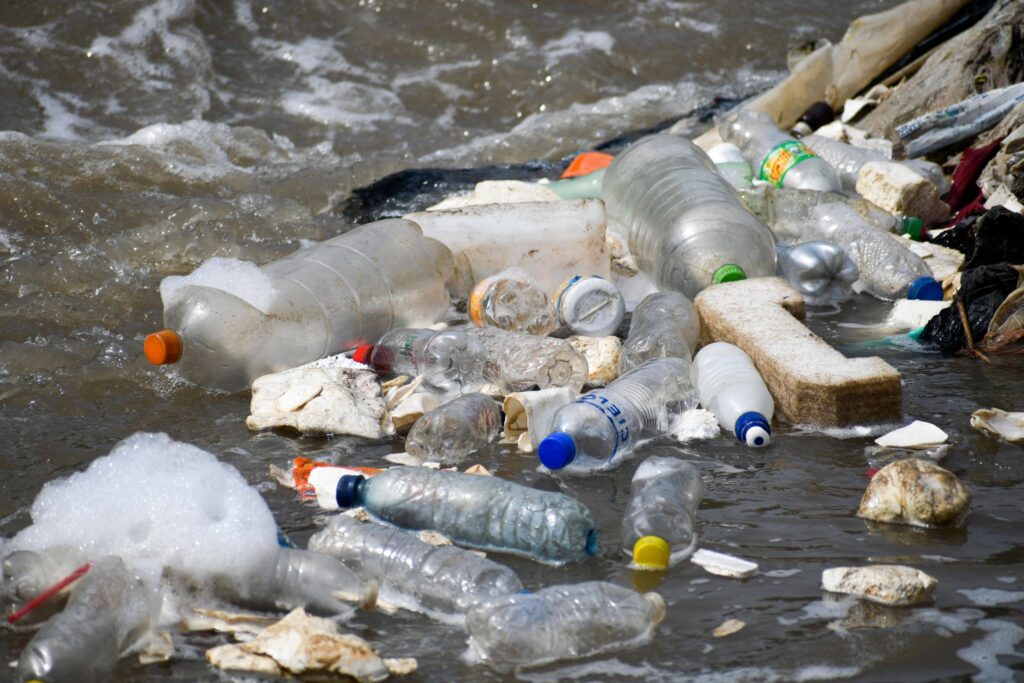

Our lakes, rivers and oceans are increasingly clogged with plastic, plus trillions of microscopic fragments thereof, from all the useful and disturbingly durable products made possible by the petroleum industry.
This deluge of waste has grown exponentially over 60 years. Some 10 million tons of bottles, nets, bags, buckets and food wrappings are deliberately or indifferently dumped each year into our waterways, where they entangle and kill marine life and damage the organs of the creatures, including, possibly, humans, that ingest them.
What can we do? For 70 years, we’ve been trying to recycle plastic, without much success. Some think it will never truly work, in large part because there are many different types of plastics and they generally can’t be recycled together. Also, recycling is still more expensive than manufacturing new plastic; the plastics industry is expected to produce three times as much plastic in 2040 as it does today.
Plastic has become so enmeshed in our ecosystem that bacteria have evolved to digest it.
Oddly enough, those bugs might now offer a ray of hope. A key barrier to cost-effective recycling is finding chemical enzymes able to break down plastic quickly, recovering the molecules originally used to make it, a crucial initial step in reforming and reusing the material.
By studying these plastic-eating bacteria, scientists have discovered some enzymes able to break down plastics far faster than was possible a decade ago. That’s a big advance from traditional recycling, which uses heat to melt plastic, leading to degraded and less useful material. Having demonstrated the new technique, a company in France called Carbios expects to soon be recycling 50,000 tons of plastic each year.
But this is likely just a beginning. The greater hope for big breakthroughs in recycling chemistry comes from our current spectacular ignorance of the microbiology of the seas and the genomics and computing technology now gearing up to change that.
We know very little about the world’s microbes. When biologists study the genetic content of seawater samples, two-thirds of what they find is unlike anything from known organisms. A recent study by researchers at the Institute of Microbiology at ETH Zurich, for example, used computational genomics to analyze more than 1,000 samples of seawater from many locations and depths and ended up producing the full genomes of some 26,000 organisms, 2,700 of which were previously unknown. (We also don’t know much about microbes in the soil. Some 99% of the genes identified in random samples of topsoil aren’t found in databases of known microbial genes.)
The microbiome — the universe of all microbial organisms — is a treasure chest of chemical leads about possible new drugs and other potentially useful biochemical compounds. The ETH Zurich study alone found more than 40,000 new biosynthetic gene clusters — biologists’ term for small clusters of associated genes that together help produce some particular bioactive molecule. For scientists, these are prime candidates in the search for new and useful pharmaceutical compounds.
Such studies are also helping scientists identify new enzymes able to digest plastics. In a study published last year, biologist Aleksej Zelezniak of Chalmers University of Technology in Sweden and colleagues identified plastic-degrading enzymes in the genomes of many bacteria, including those in the ocean and in soils. Among the ocean bacteria, they also noted a strong correlation between the diversity of such enzymes and the amount of local plastic pollution. In the brief 60 years that plastics pollution has been with us, bacteria already have responded by evolving a biochemistry to digest plastic as a food source.
In turn, this bacterial engineering offers clues about how we might produce better enzymes for recycling. From this bacterial data, using modern genomics methods and machine learning, the researchers were also able to identify more than 30,000 new candidate molecules expected to have powerful plastic-digestive properties for at least 10 different types of plastic.
There are, of course, other barriers to plastic recycling, not least a broad lack of public engagement. That’s beginning to change, spurred in part by China’s ban on importing plastic waste, which has made it harder for Western nations to hide their plastic pollution by shipping it far away.
But we need much stronger commitments from governments to tax plastic packaging and encourage packing alternatives that use less plastic, or don’t use any at all. If we take these steps, it isn’t too crazy to hope that, in 10 or 20 years, armed with more knowledge of the marine microbiome, scientists may find their way to a set of enzymes able to rapidly digest the many kinds of plastics industry might produce. If that happens, there could be hope for the oceans after all.
Recent Posts
- Astronomers detect first direct image of black hole expelling a powerful jet
- WhatsApp rolling out ‘reply with message’ feature within call notifications
- Multi-Device Pairing May Be Arriving for Apple Watch this Year
- Artificial Intelligence Discovers Hidden Giant, a Planet 5 Times Larger Than Jupiter
- Google CEO Sundar Pichai Talks Bard & The Future Of Search
Recent Comments

Astronomers detect first direct image of black hole expelling a powerful jet
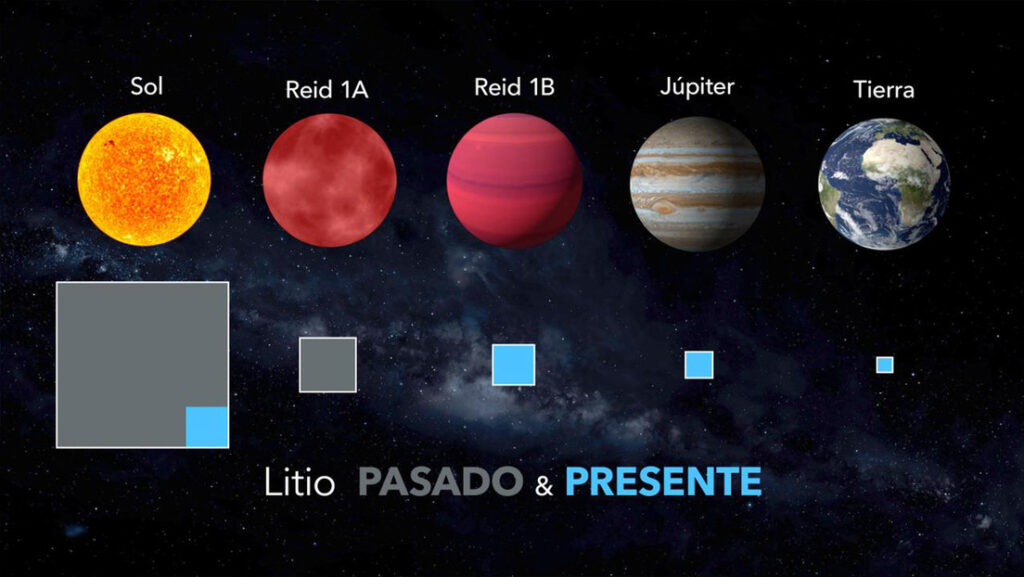
Artificial Intelligence Discovers Hidden Giant, a Planet 5 Times Larger Than Jupiter
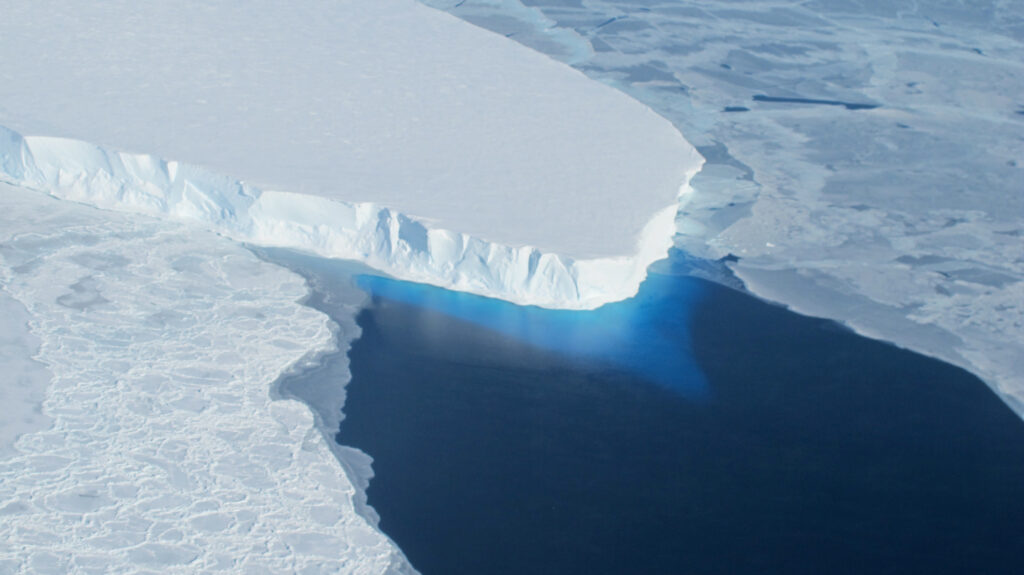

Scientists explain melting of Antarctic ice sheet dating back 9,000 years

An Unexpected Discovery: Hubble, ESA's Gaia Spot Double Quasar That Existed Over 10 Billion Years Ago

Astronomers detect first direct image of black hole expelling a powerful jet

WhatsApp rolling out ‘reply with message’ feature within call notifications

Multi-Device Pairing May Be Arriving for Apple Watch this Year


